
A new device created at the University of Notre Dame employs an innovative method for “listening in” on cells’ conversations.
Scientists have long known that RNA (ribonucleic acid) acts as a messenger inside cells, translating DNA information to help cells make proteins.
But recently, scientists have discovered that certain types of RNA venture outside the cell wall. Each of these strands of “extracellular RNA,” or exRNA, rests inside a tiny carrier “bottle” and flows along bodily fluids like a microscopic message in a bottle, carrying information to other cells.
The new appreciation for exRNA also raised a tantalizing possibility: Could we use exRNA as a way of “listening in” on cells’ conversations?
“These extracellular RNAs are a goldmine of information,” said Hsueh-Chia Chang, the Bayer Professor of Chemical and Biomolecular Engineering at the University of Notre Dame. “They can carry the early warning signs of cancer, heart disease, HIV and other life-threatening conditions.”
Chang, an expert in nanofluidics, explains that diagnosing a disease using exRNA could prove not only more effective but also faster and cheaper than existing methods, since there is enough exRNA in a small sample of blood or another bodily fluid to signal the presence of many diseases.
But intercepting and interpreting exRNA messages has been a difficult challenge. Many labs have attempted to filter them from samples of blood or other bodily fluids. Many others have used advanced centrifuges to isolate exRNA. These methods have met with little success for a simple reason: The different types of “bottles” that carry exRNA messages overlap in size and weight. Even the most advanced filters and centrifuges leave many carriers jumbled together. Labs using these methods have to add additional steps in which they add chemicals or small magnetic particles to further sort the carriers into discrete groups.
Four years ago, Chang and a team of researchers at Notre Dame decided to try a radically new approach, and their idea received support from the Common Fund of the National Institutes of Health, which selects promising “high-risk, innovative endeavors with the potential for extraordinary impact.”
Chang was joined by three other Notre Dame faculty members: Crislyn D’Souza-Schorey, the Morris Pollard Professor of Biological Sciences; David Go, vice president and associate provost for academic strategy and the Viola D. Hank Professor of Aerospace and Mechanical Engineering; and Satyajyoti Senapati, research associate professor in the Department of Chemical and Biomolecular Engineering. Postdoctoral fellow Himani Sharma served as project lead, and chemical and biomolecular engineering graduate student Vivek Yadav helped conduct the research.
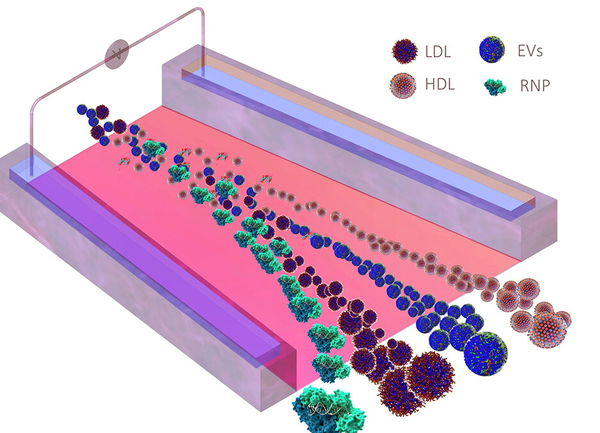
In a study published in ACS Nano, Sharma, Chang and their colleagues describe the groundbreaking device that resulted from their research. The new technology uses a combination of pH (acidity/basicity) and electrical charge to separate the carriers. The idea relies on the fact that although the carriers overlap in size and weight, each type has a distinct “isoelectric point” — the pH, or level of acidity/basicity, at which it has no positive or negative charge.
The device integrates several existing technologies developed at Notre Dame and fits neatly in the palm of the hand.
Flowing through the middle of the device is what looks like a simple stream of water. But there is something special about the stream that is not visible to the naked eye. At the left side, the water is highly acidic, with a pH about the same as a glass of grapefruit juice. On the other side of the stream, the water is highly basic, with a pH similar to a bottle of ammonia.
A special feature of the device is not just the fact that it has a pH gradient in the stream but also how it achieves that gradient. The technology is able to generate the gradient without the addition of any chemicals, making it cheaper, more eco-friendly and more efficient to run than designs that rely on added acids and bases.

The gradient comes not from a chemical but from a two-sided membrane powered by a specially designed chip. The membrane splits the water in two ions (H+ and OH-) and adds a different kind of ion to each side of the stream. One side of the membrane releases acidic hydronium ions, and the other size releases basic hydroxide ions. When the basic and acidic streams flow together, they create a pH gradient just as hot and cold streams flowing together would form hot and cold sides with a gradient of temperature through the middle of the stream. The team used the two devices running in parallel to select the pH range required for carrier separation and optimized the process using machine learning.
The pH gradient achieved what filters and centrifuges could not: It caused the exRNA carriers floating in the stream to sort themselves like colors of light passing through a prism. The different types of carriers formed lines along their isoelectric points where they could easily flow out into separate outlets.
Thanks to the new method, the research team was able to generate very pure samples (up to 97 percent pure) using less than a milliliter of blood plasma, saliva or urine. The process was also lightning-fast compared to current methods. Whereas the best existing technologies take about a day to achieve separation, the Notre Dame team was able to comprehensively sort their sample in just half an hour.
“We have filed for a patent and soon hope that the technology will be commercialized, so that it can help improve diagnoses of cancer and other diseases,” said Sharma, who won several awards for her work on the study from Notre Dame’s Harper Cancer Research Institute.
“Noncommunicable diseases are responsible for more than 70 percent of deaths worldwide, and cardiovascular disease and cancer are responsible for most of that number,” Sharma said. “Our technology shows a path to improving the way clinicians diagnose these diseases, and that could save a tremendous number of lives.”
Contact: Brandi Wampler, associate director of media relations, 574-631-2632, brandiwampler@nd.edu
Originally published by at research.nd.edu on Oct. 18.